Sanjay Iyer

Developed Artificially Intelligent (AI) workflows that leverage Large Language Models (LLMs) to guide users and streamline chemical analyses, integrating Python scripts for process automation alongside evolutionary and Bayesian optimization algorithms for efficiently exploring complex experimental search spaces.
Currently
Finishing Ph.D. at Purdue University (Chopra Lab). Exploring opportunities starting in 2026 (analytical, computational, or AI‑driven R&D).
Selected Projects
CLAW‑MRM
End‑to‑end automation for lipidomics MRM: worklist generation, QC agents, parsing, stats, and visualization.
- Automates lipid profiling—from parsing raw binary (.d) files into organized pandas DataFrames to annotation using custom MRM transition databases and robust statistical analysis.
- Applies trimmed mean of M-values (TMM) normalization for lipid load correction and generalized linear models (GLMs) in edgeR for overdispersed count data analysis, providing a robust statistical framework for lipidomics.
- Integrates LIGER (lipidome gene enrichment reactions) to connect lipid expression with gene activation and suppression patterns, highlighting biologically relevant pathways.
- Incorporates a language user interface (LUI) with interactive artificially intelligent (AI) agents, providing a conversational way to perform statistical and bioinformatics workflows.
- Applied this pipeline to Alzheimer’s disease mouse models, profiling nearly 1,500 MRM transitions across 11 lipid classes, and revealing metabolic pathways enriched in differentially expressed lipids.
Paper · Code · Project Page
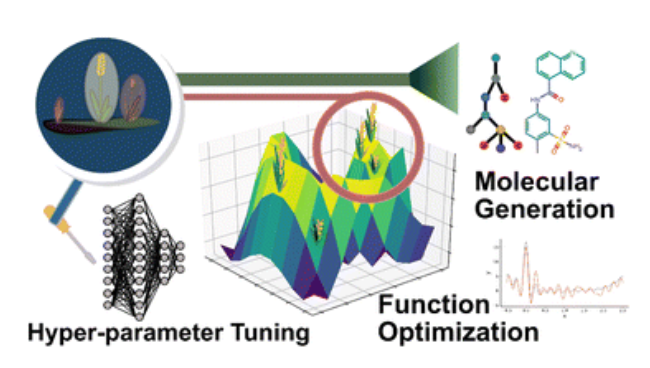
Paddy: Automated Data Validation
Paddy is a versatile, biologically inspired evolutionary algorithm designed to efficiently optimize complex chemical systems while avoiding local minima.
- Robust and adaptable: Demonstrates strong performance across diverse benchmarks, from mathematical functions to chemical optimization tasks.
- Exploratory sampling: Effectively avoids early convergence by bypassing local optima, ensuring global solutions are identified.
- Chemical applications: Enables hyperparameter tuning, molecular generation, and experimental design, accelerating automated experimentation and discovery.
Paper · Code · Project Page
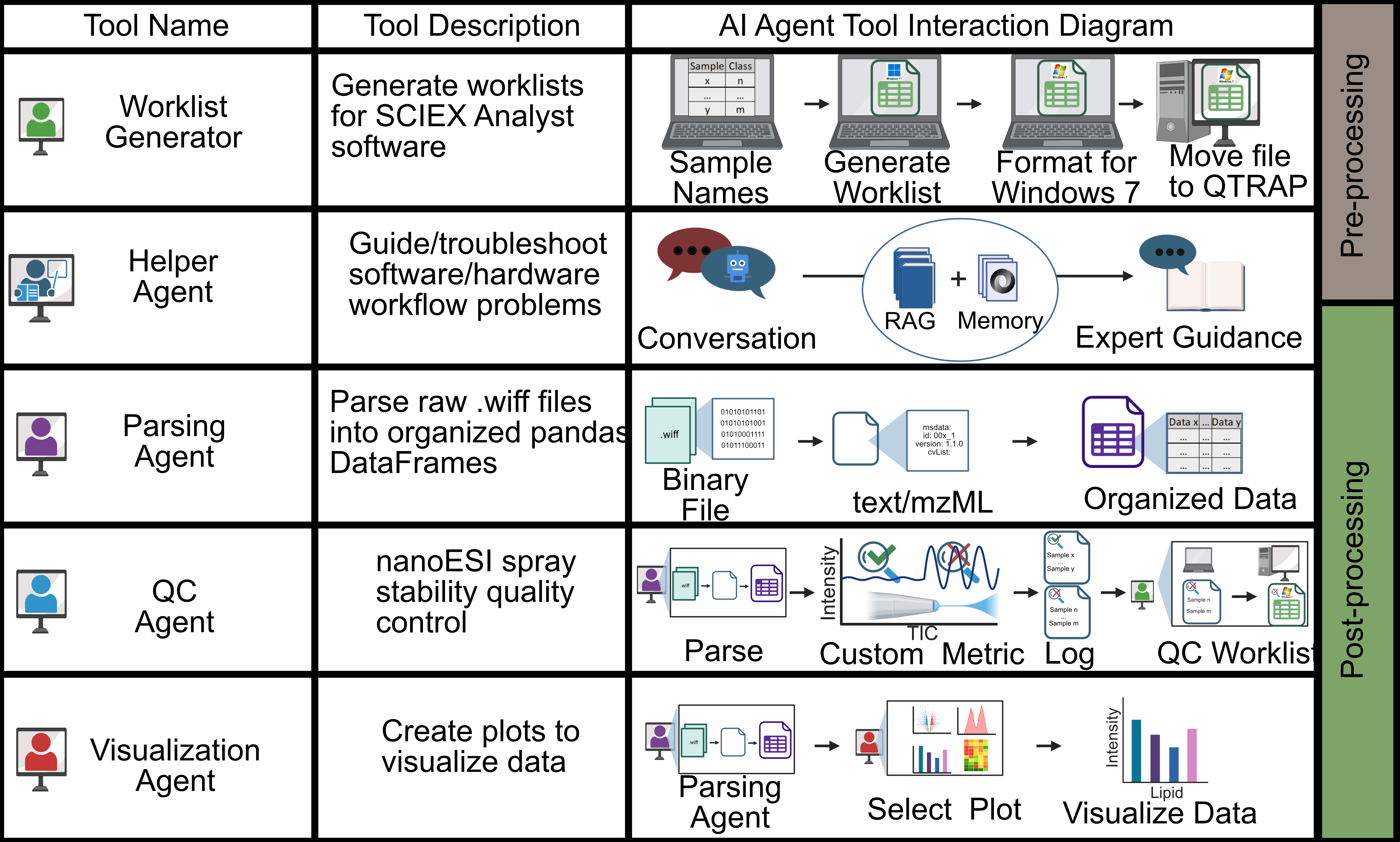
Automated Agentic Chip-based nanoESI-MS/MS
An LLM-driven AI agent framework for chip-based nanoESI-MS/MS automates experimental and analytical tasks while remaining accessible, adaptable, and informed by both literature and lab-specific knowledge. Manuscript in preparation.
- AI-driven automation: Integrates LLM reasoning with tool-based execution to autonomously handle worklist creation, QC, data conversion, visualization, and literature retrieval.
- Accessible workflows: Eliminates coding barriers, enabling non-specialists to run end-to-end chip-based nanoESI-MS/MS experiments independently.
- Adaptive and evolving framework: Modular design preserves lab knowledge, integrates literature, and flexibly updates with new methods and technologies.
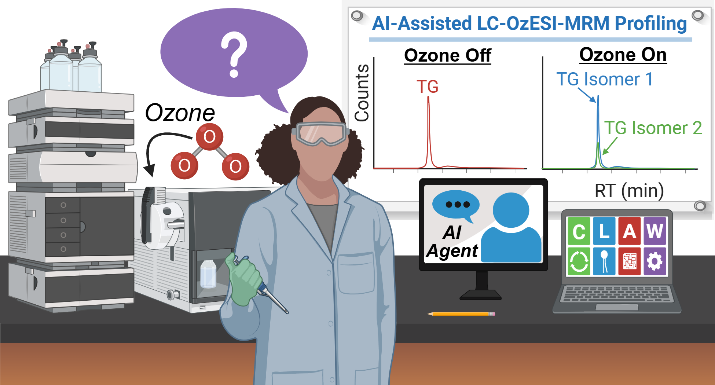
OzESI‑MRM
AI-powered LC-OzESI-MRM enables rapid, accessible, and isomer-specific fatty acid profiling to drive biomarker discovery and precision lipidomics.
- AI-powered LC-OzESI-MRM platform: Combines ozone-induced fragmentation with automated informatics and AI agents to achieve high-throughput, isomer-specific fatty acid profiling.
- Broad accessibility and biological relevance: Demonstrated on standard FA mixtures, NIST 1950 plasma, and disease models (Alzheimer’s, muscular dystrophy), uncovering extensive isomer diversity and disease-associated alterations.
- Improved resolution and reproducibility: Delivers rapid, robust, and accurate FA characterization beyond traditional methods, accelerating biomarker discovery and enabling precision lipidomics for therapeutic development.
Publications
Contact
Contact: SanjayIyer01@gmail.com iyer95@purdue.edu
For consulting or collaborations, include a brief summary and timeline.